Electron transport chain
An electron transport chain (ETC[1]) is a series of protein complexes and other molecules which transfer electrons from electron donors to electron acceptors via redox reactions (both reduction and oxidation occurring simultaneously) and couples this electron transfer with the transfer of protons (H+ ions) across a membrane. Many of the enzymes in the electron transport chain are embedded within the membrane.
The flow of electrons through the electron transport chain is an exergonic process. The energy from the redox reactions creates an electrochemical proton gradient that drives the synthesis of adenosine triphosphate (ATP). In aerobic respiration, the flow of electrons terminates with molecular oxygen as the final electron acceptor. In anaerobic respiration, other electron acceptors are used, such as sulfate.
In an electron transport chain, the redox reactions are driven by the difference in the Gibbs free energy of reactants and products. The free energy released when a higher-energy electron donor and acceptor convert to lower-energy products, while electrons are transferred from a lower to a higher redox potential, is used by the complexes in the electron transport chain to create an electrochemical gradient of ions. It is this electrochemical gradient that drives the synthesis of ATP via coupling with oxidative phosphorylation with ATP synthase.[2]
In eukaryotic organisms, the electron transport chain, and site of oxidative phosphorylation, is found on the inner mitochondrial membrane. The energy released by reactions of oxygen and reduced compounds such as cytochrome c and (indirectly) NADH and FADH2 is used by the electron transport chain to pump protons into the intermembrane space, generating the electrochemical gradient over the inner mitochondrial membrane. In photosynthetic eukaryotes, the electron transport chain is found on the thylakoid membrane. Here, light energy drives electron transport through a proton pump and the resulting proton gradient causes subsequent synthesis of ATP. In bacteria, the electron transport chain can vary between species but it always constitutes a set of redox reactions that are coupled to the synthesis of ATP through the generation of an electrochemical gradient and oxidative phosphorylation through ATP synthase.[3]
Mitochondrial electron transport chains
[edit]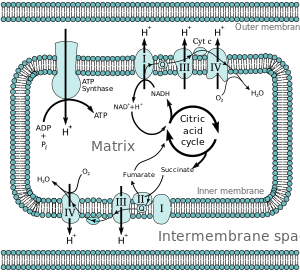
Most eukaryotic cells have mitochondria, which produce ATP from reactions of oxygen with products of the citric acid cycle, fatty acid metabolism, and amino acid metabolism. At the inner mitochondrial membrane, electrons from NADH and FADH2 pass through the electron transport chain to oxygen, which provides the energy driving the process as it is reduced to water.[4] The electron transport chain comprises an enzymatic series of electron donors and acceptors. Each electron donor will pass electrons to an acceptor of higher redox potential, which in turn donates these electrons to another acceptor, a process that continues down the series until electrons are passed to oxygen, the terminal electron acceptor in the chain. Each reaction releases energy because a higher-energy donor and acceptor convert to lower-energy products. Via the transferred electrons, this energy is used to generate a proton gradient across the mitochondrial membrane by "pumping" protons into the intermembrane space, producing a state of higher free energy that has the potential to do work. This entire process is called oxidative phosphorylation since ADP is phosphorylated to ATP by using the electrochemical gradient that the redox reactions of the electron transport chain have established driven by energy-releasing reactions of oxygen.
Mitochondrial redox carriers
[edit]Energy associated with the transfer of electrons down the electron transport chain is used to pump protons from the mitochondrial matrix into the intermembrane space, creating an electrochemical proton gradient (ΔpH) across the inner mitochondrial membrane. This proton gradient is largely but not exclusively responsible for the mitochondrial membrane potential (ΔΨM).[5] It allows ATP synthase to use the flow of H+ through the enzyme back into the matrix to generate ATP from adenosine diphosphate (ADP) and inorganic phosphate. Complex I (NADH coenzyme Q reductase; labeled I) accepts electrons from the Krebs cycle electron carrier nicotinamide adenine dinucleotide (NADH), and passes them to coenzyme Q (ubiquinone; labeled Q), which also receives electrons from Complex II (succinate dehydrogenase; labeled II). Q passes electrons to Complex III (cytochrome bc1 complex; labeled III), which passes them to cytochrome c (cyt c). Cyt c passes electrons to Complex IV (cytochrome c oxidase; labeled IV).
Four membrane-bound complexes have been identified in mitochondria. Each is an extremely complex transmembrane structure that is embedded in the inner membrane. Three of them are proton pumps. The structures are electrically connected by lipid-soluble electron carriers and water-soluble electron carriers. The overall electron transport chain can be summarized as follows:
NADH, H+ → Complex I → Q → Complex III → cytochrome c → Complex IV → H2O
↑
Complex II
↑
Succinate
Complex I
[edit]In Complex I (NADH ubiquinone oxidoreductase, Type I NADH dehydrogenase, or mitochondrial complex I; EC 1.6.5.3), two electrons are removed from NADH and transferred to a lipid-soluble carrier, ubiquinone (Q). The reduced product, ubiquinol (QH2), freely diffuses within the membrane, and Complex I translocates four protons (H+) across the membrane, thus producing a proton gradient. Complex I is one of the main sites at which premature electron leakage to oxygen occurs, thus being one of the main sites of production of superoxide.[6]
The pathway of electrons is as follows:
NADH is oxidized to NAD+, by reducing flavin mononucleotide to FMNH2 in one two-electron step. FMNH2 is then oxidized in two one-electron steps, through a semiquinone intermediate. Each electron thus transfers from the FMNH2 to an Fe–S cluster, from the Fe-S cluster to ubiquinone (Q). Transfer of the first electron results in the free-radical (semiquinone) form of Q, and transfer of the second electron reduces the semiquinone form to the ubiquinol form, QH2. During this process, four protons are translocated from the mitochondrial matrix to the intermembrane space.[7] As the electrons move through the complex an electron current is produced along the 180 Angstrom width of the complex within the membrane. This current powers the active transport of four protons to the intermembrane space per two electrons from NADH.[8]
Complex II
[edit]In Complex II (succinate dehydrogenase or succinate-CoQ reductase; EC 1.3.5.1) additional electrons are delivered into the quinone pool (Q) originating from succinate and transferred (via flavin adenine dinucleotide (FAD)) to Q. Complex II consists of four protein subunits: succinate dehydrogenase (SDHA); succinate dehydrogenase [ubiquinone] iron–sulfur subunit mitochondrial (SDHB); succinate dehydrogenase complex subunit C (SDHC); and succinate dehydrogenase complex subunit D (SDHD). Other electron donors (e.g., fatty acids and glycerol 3-phosphate) also direct electrons into Q (via FAD). Complex II is a parallel electron transport pathway to Complex I, but unlike Complex I, no protons are transported to the intermembrane space in this pathway. Therefore, the pathway through Complex II contributes less energy to the overall electron transport chain process.
Complex III
[edit]In Complex III (cytochrome bc1 complex or CoQH2-cytochrome c reductase; EC 1.10.2.2), the Q-cycle contributes to the proton gradient by an asymmetric absorption/release of protons. Two electrons are removed from QH2 at the QO site and sequentially transferred to two molecules of cytochrome c, a water-soluble electron carrier located within the intermembrane space. The two other electrons sequentially pass across the protein to the Qi site where the quinone part of ubiquinone is reduced to quinol. A proton gradient is formed by one quinol () oxidations at the Qo site to form one quinone () at the Qi site. (In total, four protons are translocated: two protons reduce quinone to quinol and two protons are released from two ubiquinol molecules.)
When electron transfer is reduced (by a high membrane potential or respiratory inhibitors such as antimycin A), Complex III may leak electrons to molecular oxygen, resulting in superoxide formation.
This complex is inhibited by dimercaprol (British Anti-Lewisite, BAL), naphthoquinone and antimycin.
Complex IV
[edit]In Complex IV (cytochrome c oxidase; EC 1.9.3.1), sometimes called cytochrome AA3, four electrons are removed from four molecules of cytochrome c and transferred to molecular oxygen (O2) and four protons, producing two molecules of water. The complex contains coordinated copper ions and several heme groups. At the same time, eight protons are removed from the mitochondrial matrix (although only four are translocated across the membrane), contributing to the proton gradient. The exact details of proton pumping in Complex IV are still under study.[9] Cyanide is an inhibitor of Complex IV.
Coupling with oxidative phosphorylation
[edit]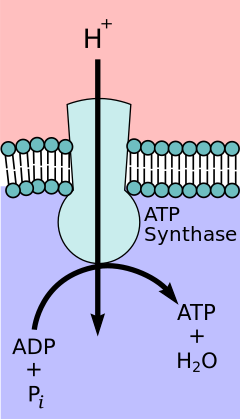
According to the chemiosmotic coupling hypothesis, proposed by Nobel Prize in Chemistry winner Peter D. Mitchell, the electron transport chain and oxidative phosphorylation are coupled by a proton gradient across the inner mitochondrial membrane. The efflux of protons from the mitochondrial matrix creates an electrochemical gradient (proton gradient). This gradient is used by the FOF1 ATP synthase complex to make ATP via oxidative phosphorylation. ATP synthase is sometimes described as Complex V of the electron transport chain.[10] The FO component of ATP synthase acts as an ion channel that provides for a proton flux back into the mitochondrial matrix. It is composed of a, b and c subunits. Protons in the inter-membrane space of mitochondria first enter the ATP synthase complex through an a subunit channel. Then protons move to the c subunits.[11] The number of c subunits determines how many protons are required to make the FO turn one full revolution. For example, in humans, there are 8 c subunits, thus 8 protons are required.[12] After c subunits, protons finally enter the matrix through an a subunit channel that opens into the mitochondrial matrix.[11] This reflux releases free energy produced during the generation of the oxidized forms of the electron carriers (NAD+ and Q) with energy provided by O2. The free energy is used to drive ATP synthesis, catalyzed by the F1 component of the complex.[13]
Coupling with oxidative phosphorylation is a key step for ATP production. However, in specific cases, uncoupling the two processes may be biologically useful. The uncoupling protein, thermogenin—present in the inner mitochondrial membrane of brown adipose tissue—provides for an alternative flow of protons back to the inner mitochondrial matrix. Thyroxine is also a natural uncoupler. This alternative flow results in thermogenesis rather than ATP production.[14]
Reverse electron flow
[edit]Reverse electron flow is the transfer of electrons through the electron transport chain through the reverse redox reactions. Usually requiring a significant amount of energy to be used, this can reduce the oxidized forms of electron donors. For example, NAD+ can be reduced to NADH by Complex I.[15] There are several factors that have been shown to induce reverse electron flow. However, more work needs to be done to confirm this. One example is blockage of ATP synthase, resulting in a build-up of protons and therefore a higher proton-motive force, inducing reverse electron flow.[16]
Prokaryotic electron transport chains
[edit]This section needs additional citations for verification. (December 2023) |
In eukaryotes, NADH is the most important electron donor. The associated electron transport chain is NADH → Complex I → Q → Complex III → cytochrome c → Complex IV → O2 where Complexes I, III and IV are proton pumps, while Q and cytochrome c are mobile electron carriers. The electron acceptor for this process is molecular oxygen.
In prokaryotes (bacteria and archaea) the situation is more complicated, because there are several different electron donors and several different electron acceptors. The generalized electron transport chain in bacteria is:
Donor Donor Donor
↓ ↓ ↓
dehydrogenase → quinone → bc1 → cytochrome
↓ ↓
oxidase(reductase) oxidase(reductase)
↓ ↓
Acceptor Acceptor
Electrons can enter the chain at three levels: at the level of a dehydrogenase, at the level of the quinone pool, or at the level of a mobile cytochrome electron carrier. These levels correspond to successively more positive redox potentials, or to successively decreased potential differences relative to the terminal electron acceptor. In other words, they correspond to successively smaller Gibbs free energy changes for the overall redox reaction.
Individual bacteria use multiple electron transport chains, often simultaneously. Bacteria can use a number of different electron donors, a number of different dehydrogenases, a number of different oxidases and reductases, and a number of different electron acceptors. For example, E. coli (when growing aerobically using glucose and oxygen as an energy source) uses two different NADH dehydrogenases and two different quinol oxidases, for a total of four different electron transport chains operating simultaneously.
A common feature of all electron transport chains is the presence of a proton pump to create an electrochemical gradient over a membrane. Bacterial electron transport chains may contain as many as three proton pumps, like mitochondria, or they may contain two or at least one.
Electron donors
[edit]In the current biosphere, the most common electron donors are organic molecules. Organisms that use organic molecules as an electron source are called organotrophs. Chemoorganotrophs (animals, fungi, protists) and photolithotrophs (plants and algae) constitute the vast majority of all familiar life forms.
Some prokaryotes can use inorganic matter as an electron source. Such an organism is called a (chemo)lithotroph ("rock-eater"). Inorganic electron donors include hydrogen, carbon monoxide, ammonia, nitrite, sulfur, sulfide, manganese oxide, and ferrous iron. Lithotrophs have been found growing in rock formations thousands of meters below the surface of Earth. Because of their volume of distribution, lithotrophs may actually outnumber organotrophs and phototrophs in our biosphere.
The use of inorganic electron donors such as hydrogen as an energy source is of particular interest in the study of evolution. This type of metabolism must logically have preceded the use of organic molecules and oxygen as an energy source.
Dehydrogenases: equivalents to complexes I and II
[edit]Bacteria can use several different electron donors. When organic matter is the electron source, the donor may be NADH or succinate, in which case electrons enter the electron transport chain via NADH dehydrogenase (similar to Complex I in mitochondria) or succinate dehydrogenase (similar to Complex II). Other dehydrogenases may be used to process different energy sources: formate dehydrogenase, lactate dehydrogenase, glyceraldehyde-3-phosphate dehydrogenase, H2 dehydrogenase (hydrogenase), electron transport chain. Some dehydrogenases are also proton pumps, while others funnel electrons into the quinone pool. Most dehydrogenases show induced expression in the bacterial cell in response to metabolic needs triggered by the environment in which the cells grow. In the case of lactate dehydrogenase in E. coli, the enzyme is used aerobically and in combination with other dehydrogenases. It is inducible and is expressed when the concentration of DL-lactate in the cell is high.[citation needed]
Quinone carriers
[edit]Quinones are mobile, lipid-soluble carriers that shuttle electrons (and protons) between large, relatively immobile macromolecular complexes embedded in the membrane. Bacteria use ubiquinone (Coenzyme Q, the same quinone that mitochondria use) and related quinones such as menaquinone (Vitamin K2). Archaea in the genus Sulfolobus use caldariellaquinone.[17] The use of different quinones is due to slight changes in redox potentials caused by changes in structure. The change in redox potentials of these quinones may be suited to changes in the electron acceptors or variations of redox potentials in bacterial complexes.[18]
Proton pumps
[edit]A proton pump is any process that creates a proton gradient across a membrane. Protons can be physically moved across a membrane, as seen in mitochondrial Complexes I and IV. The same effect can be produced by moving electrons in the opposite direction. The result is the disappearance of a proton from the cytoplasm and the appearance of a proton in the periplasm. Mitochondrial Complex III is this second type of proton pump, which is mediated by a quinone (the Q cycle).
Some dehydrogenases are proton pumps, while others are not. Most oxidases and reductases are proton pumps, but some are not. Cytochrome bc1 is a proton pump found in many, but not all, bacteria (not in E. coli). As the name implies, bacterial bc1 is similar to mitochondrial bc1 (Complex III).
Cytochrome electron carriers
[edit]Cytochromes are proteins that contain iron. They are found in two very different environments.
Some cytochromes are water-soluble carriers that shuttle electrons to and from large, immobile macromolecular structures imbedded in the membrane. The mobile cytochrome electron carrier in mitochondria is cytochrome c. Bacteria use a number of different mobile cytochrome electron carriers.
Other cytochromes are found within macromolecules such as Complex III and Complex IV. They also function as electron carriers, but in a very different, intramolecular, solid-state environment.
Electrons may enter an electron transport chain at the level of a mobile cytochrome or quinone carrier. For example, electrons from inorganic electron donors (nitrite, ferrous iron, electron transport chain) enter the electron transport chain at the cytochrome level. When electrons enter at a redox level greater than NADH, the electron transport chain must operate in reverse to produce this necessary, higher-energy molecule.
Electron acceptors and terminal oxidase/reductase
[edit]This section may require cleanup to meet Wikipedia's quality standards. The specific problem is: We talk as if oxidases are not also reductases, and as if reductases are not also oxidizing something. That's messed up. (December 2023) |
As there are a number of different electron donors (organic matter in organotrophs, inorganic matter in lithotrophs), there are a number of different electron acceptors, both organic and inorganic. As with other steps of the ETC, an enzyme is required to help with the process.
If oxygen is available, it is most often used as the terminal electron acceptor in aerobic bacteria and facultative anaerobes. An oxidase reduces the O2 to water while oxidizing something else. In mitochondria, the terminal membrane complex (Complex IV) is cytochrome oxidase, which oxidizes the cytochrome. Aerobic bacteria use a number of differet terminal oxidases. For example, E. coli (a facultative anaerobe) does not have a cytochrome oxidase or a bc1 complex. Under aerobic conditions, it uses two different terminal quinol oxidases (both proton pumps) to reduce oxygen to water.
Bacterial terminal oxidases can be split into classes according to the molecules act as terminal electron acceptors. Class I oxidases are cytochrome oxidases and use oxygen as the terminal electron acceptor. Class II oxidases are quinol oxidases and can use a variety of terminal electron acceptors. Both of these classes can be subdivided into categories based on what redox-active components they contain. E.g. Heme aa3 Class 1 terminal oxidases are much more efficient than Class 2 terminal oxidases.[2]
Mostly in anaerobic environments different electron acceptors are used, including nitrate, nitrite, ferric iron, sulfate, carbon dioxide, and small organic molecules such as fumarate. When bacteria grow in anaerobic environments, the terminal electron acceptor is reduced by an enzyme called a reductase. E. coli can use fumarate reductase, nitrate reductase, nitrite reductase, DMSO reductase, or trimethylamine-N-oxide reductase, depending on the availability of these acceptors in the environment.
Most terminal oxidases and reductases are inducible. They are synthesized by the organism as needed, in response to specific environmental conditions.
Photosynthetic
[edit]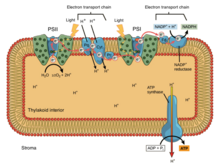
In oxidative phosphorylation, electrons are transferred from an electron donor such as NADH to an acceptor such as O2 through an electron transport chain, releasing energy. In photophosphorylation, the energy of sunlight is used to create a high-energy electron donor which can subsequently reduce oxidized components and couple to ATP synthesis via proton translocation by the electron transport chain.[9]
Photosynthetic electron transport chains, like the mitochondrial chain, can be considered as a special case of the bacterial systems. They use mobile, lipid-soluble quinone carriers (phylloquinone and plastoquinone) and mobile, water-soluble carriers (cytochromes). They also contain a proton pump. The proton pump in all photosynthetic chains resembles mitochondrial Complex III. The commonly-held theory of symbiogenesis proposes that both organelles descended from bacteria.
See also
[edit]- Charge-transfer complex
- CoRR hypothesis
- Electron equivalent
- Hydrogen hypothesis
- Respirasome
- Electric bacteria
References
[edit]- ^ Lyall, Fiona (2010). "Biochemistry". Basic Science in Obstetrics and Gynaecology. pp. 143–171. doi:10.1016/B978-0-443-10281-3.00013-0. ISBN 978-0-443-10281-3.
- ^ a b Anraku Y (June 1988). "Bacterial electron transport chains". Annual Review of Biochemistry. 57 (1): 101–32. doi:10.1146/annurev.bi.57.070188.000533. PMID 3052268.
- ^ Kracke F, Vassilev I, Krömer JO (2015). "Microbial electron transport and energy conservation - the foundation for optimizing bioelectrochemical systems". Frontiers in Microbiology. 6: 575. doi:10.3389/fmicb.2015.00575. PMC 4463002. PMID 26124754. – This source shows four ETCs (Geobacter, Shewanella, Moorella , Acetobacterium) in figures 1 and 2.
- ^ Waldenström JG (2009-04-24). "Biochemistry. By Lubert Stryer". Acta Medica Scandinavica. 198 (1–6): 436. doi:10.1111/j.0954-6820.1975.tb19571.x. ISSN 0001-6101.
- ^ Zorova LD, Popkov VA, Plotnikov EY, Silachev DN, Pevzner IB, Jankauskas SS, et al. (July 2018). "Mitochondrial membrane potential". Analytical Biochemistry. 552: 50–59. doi:10.1016/j.ab.2017.07.009. PMC 5792320. PMID 28711444.
- ^ Lauren, Biochemistry, Johnson/Cole, 2010, pp 598-611
- ^ Garrett & Grisham, Biochemistry, Brooks/Cole, 2010, pp 598-611
- ^ Garrett R, Grisham CM (2016). biochemistry. Boston: Cengage. p. 687. ISBN 978-1-305-57720-6.
- ^ a b Stryer. Biochemistry. toppan. OCLC 785100491.
- ^ Jonckheere AI, Smeitink JA, Rodenburg RJ (March 2012). "Mitochondrial ATP synthase: architecture, function and pathology". Journal of Inherited Metabolic Disease. 35 (2): 211–25. doi:10.1007/s10545-011-9382-9. PMC 3278611. PMID 21874297.
- ^ a b Garrett RH, Grisham CM (2012). Biochemistry (5th ed.). Cengage learning. p. 664. ISBN 978-1-133-10629-6.
- ^ Fillingame RH, Angevine CM, Dmitriev OY (November 2003). "Mechanics of coupling proton movements to c-ring rotation in ATP synthase". FEBS Letters. 555 (1): 29–34. doi:10.1016/S0014-5793(03)01101-3. PMID 14630314. S2CID 38896804.
- ^ Berg JM, Tymoczko JL, Stryer L (2002-01-01). "A Proton Gradient Powers the Synthesis of ATP".
{{cite journal}}: Cite journal requires|journal=(help) - ^ Cannon B, Nedergaard J (January 2004). "Brown adipose tissue: function and physiological significance". Physiological Reviews. 84 (1): 277–359. doi:10.1152/physrev.00015.2003. PMID 14715917.
- ^ Kim BH, Gadd GM (2008). "Introduction to bacterial physiology and metabolism". Bacterial Physiology and Metabolism. Cambridge University Press. pp. 1–6. doi:10.1017/cbo9780511790461.002. ISBN 978-0-511-79046-1.
- ^ Mills EL, Kelly B, Logan A, Costa AS, Varma M, Bryant CE, et al. (October 2016). "Succinate Dehydrogenase Supports Metabolic Repurposing of Mitochondria to Drive Inflammatory Macrophages". Cell. 167 (2): 457–470.e13. doi:10.1016/j.cell.2016.08.064. PMC 5863951. PMID 27667687.
- ^ EC 1.3.5.1
- ^ Ingledew WJ, Poole RK (September 1984). "The respiratory chains of Escherichia coli". Microbiological Reviews. 48 (3): 222–71. doi:10.1128/mmbr.48.3.222-271.1984. PMC 373010. PMID 6387427.
Further reading
[edit]- Fenchel T, King GM, Blackburn TH (September 2006). Bacterial Biogeochemistry: The Ecophysiology of Mineral Cycling (2nd ed.). Elsevier. ISBN 978-0-12-103455-9.
- Lengeler JW (January 1999). Drews G; Schlegel HG (eds.). Biology of the Prokaryotes. Blackwell Science. ISBN 978-0-632-05357-5.
- Nelson DL, Cox MM (April 2005). Lehninger Principles of Biochemistry (4th ed.). W. H. Freeman. ISBN 978-0-7167-4339-2.
- Nicholls DG, Ferguson SJ (July 2002). Bioenergetics 3. Academic Press. ISBN 978-0-12-518121-1.
- Stumm W; Morgan JJ (1996). Aquatic Chemistry (3rd ed.). John Wiley & Sons. ISBN 978-0-471-51185-4.
- Thauer RK, Jungermann K, Decker K (March 1977). "Energy conservation in chemotrophic anaerobic bacteria". Bacteriological Reviews. 41 (1): 100–80. doi:10.1128/MMBR.41.1.100-180.1977. PMC 413997. PMID 860983.
- White D (September 1999). The Physiology and Biochemistry of Prokaryotes (2nd ed.). Oxford University Press. ISBN 978-0-19-512579-5.
- Voet D, Voet JG (March 2004). Biochemistry. Vol. 28 (3rd ed.). John Wiley & Sons. pp. 124. doi:10.1016/s0307-4412(00)00032-7. ISBN 978-0-471-58651-7. PMID 10878303.
{{cite book}}:|journal=ignored (help) - Kim HS, Patel K, Muldoon-Jacobs K, Bisht KS, Aykin-Burns N, Pennington JD, et al. (January 2010). "SIRT3 is a mitochondria-localized tumor suppressor required for maintenance of mitochondrial integrity and metabolism during stress". Cancer Cell. 17 (1): 41–52. doi:10.1016/j.ccr.2009.11.023. PMC 3711519. PMID 20129246.
- Raimondi V, Ciccarese F, Ciminale V (January 2020). "Oncogenic pathways and the electron transport chain: a dangeROS liaison". Br J Cancer. 122 (2): 168–181. doi:10.1038/s41416-019-0651-y. PMC 7052168. PMID 31819197.
- Reguera, Gemma (29 May 2018). "Biological electron transport goes the extra mile". Proceedings of the National Academy of Sciences. 115 (22): 5632–5634. Bibcode:2018PNAS..115.5632R. doi:10.1073/pnas.1806580115. PMC 5984551. PMID 29769327. – Editorial commentary mentioning two unusual ETCs: that of Geobacter sulfurreducens and that of cable bacteria. Also has schematic of E. coli ETC.
External links
[edit]- Electron+Transport+Chain+Complex+Proteins at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- Khan Academy, video lecture
- KEGG pathway: Oxidative phosphorylation, overlaid with genes found in Pseudomonas fluorescens Pf0-1. Click "help" for a how-to.








